Large proteins – large differences
22.06.2020
 BIOTEC
BIOTEC
International cancer genome sequencing projects discovered that the sister genes, MLL3 and MLL4, are mutated in almost every type of cancer at high frequencies. Despite their remarkably prominent association with malignant diseases, the essential roles of these genes in mammalian development has not been determined until now. Both genes originate from a gene duplication and are also related to the major leukemia gene, MLL1 and it’s sister MLL2. These genes encode epigenetic regulators involved in the maintenance of transcriptional activity. MLL3 and MLL4 are huge proteins, amongst the largest proteins described and easily the largest proteins found in the mammalian nucleus.
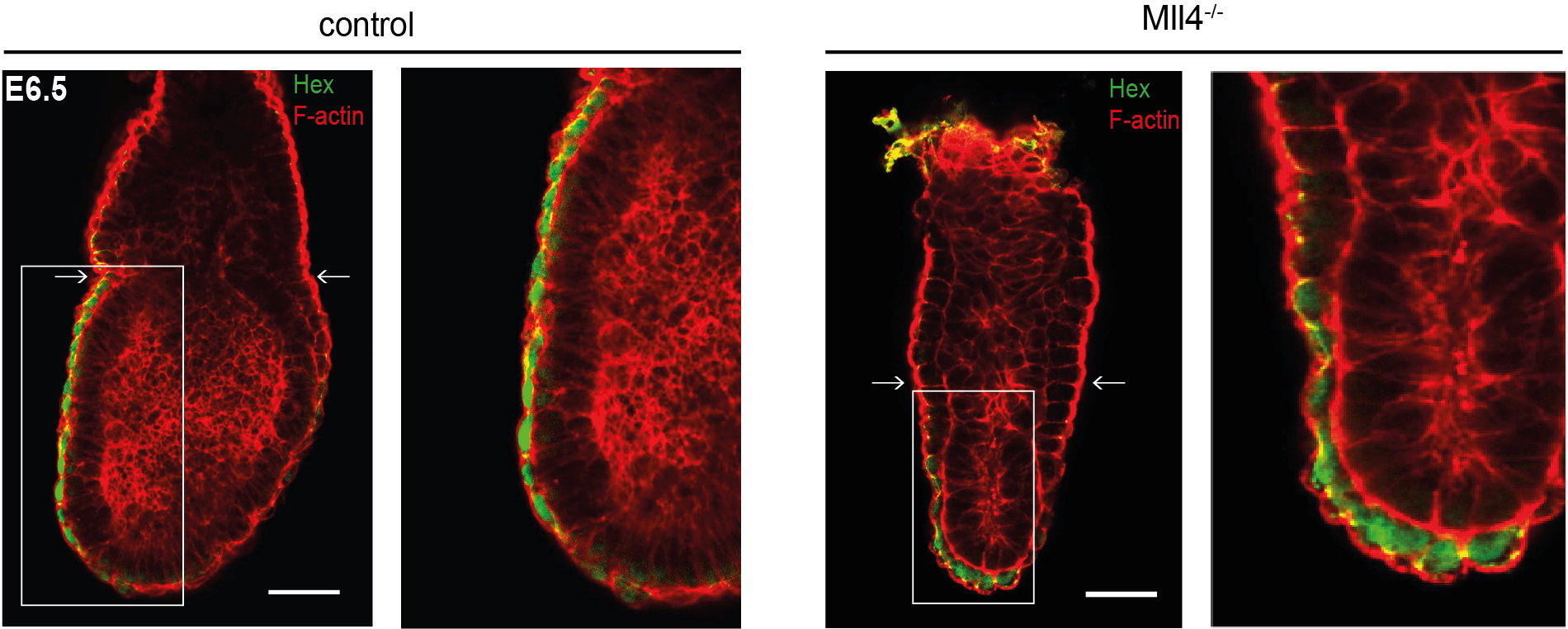
As now published in Development, the team led by Dr. Andrea Kranz and Prof. Francis Stewart report that both proteins play very different and very specific essential roles in mouse development. MLL3 is only essential for the final maturation of the lung at the very end of fetal development, which is required for the first breath of the newborn pup, whereas MLL4 is responsible for the formation of the second embryonic axis at the very beginning of mouse development. MLL4 is required for the cuboidal to squamous transition of the Anterior Visceral Endoderm (AVE) that precedes AVE migration to define the anterior to posterior embryonic axis.
“These are unexpected findings”, explains Dr. Kranz. “The extraordinary prevalence of MLL3 and MLL4 mutations in cancer implies that both genes play vital roles in most cell types. However, our knockout analysis identified an extremely precise role in just one specific cell type each. This is not consistent with their roles as general transcription factors as expected from other studies of epigenetic regulators. Together with the highly specific roles played by MLL1 and MLL2 revealed by our earlier studies, a thorough reappraisal of these important epigenetic regulators and their association to cancer development is required”.
The study completes a series of mouse knockouts to examine epigenetic regulators in mouse development by the Stewart and Anastassiadis labs at TU Dresden BIOTEC, Center for Molecular and Cellular Bioengineering (CMCB), in cooperation with the Max Planck Institute of Molecular Cell Biology and Genetics in Dresden and the DRESDEN-concept Genome Center. It was financed by funds from the Else Kröner-Fresenius-Stiftung, the Deutsche Forschungsgemeinschaft, the Deutsche Krebshilfe and the Scholarship Program for the Promotion of Early-Career Female Scientists of Technische Universität Dresden.